Protein Structure, Membrane Protein Production, & Structure-Guided Drug Design
Mistic
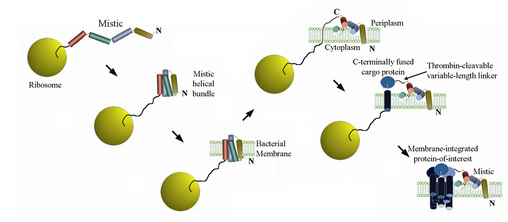
Our laboratory focuses on problems related to carcinogenesis in protein structure, biochemistry and biophysics with an emphasis on membrane proteins. Our understanding of membrane proteins has significantly lagged that of their soluble counterparts due to the numerous difficulties that arise when trying to analyze this class of proteins with conventional biophysical techniques. Arguably, the most acute of these obstacles is in fact simply producing these proteins in sufficient quantities for structural analysis. Most human and, more generally, eukaryotic membrane proteins cannot be expressed in traditional recombinant systems. Our work has led to the discovery of a protein named Mistic that circumvents this primary bottleneck. When fused to a target protein of interest, Mistic acts analogously to a “super-signal sequence,” facilitating the expression of a variety of membrane proteins in their membrane-integrated conformations within the lipid bilayer of recombinant E. coli bacteria. We continue to develop this technology to extend its application to an increasingly diverse array of proteins by surmounting technical difficulties that exist due to steric and topological restrictions imposed by the planar nature of the lipid membrane.
Mitochondrial membrane-associated apoptosis regulators
Mistic is having a major impact on the field of membrane protein structural biology allowing the study of certain scientific problems that were until recently intractable. We are applying this technology to the study of human mitochondrial proteins involved in the regulation of apoptosis. The committing step toward initiation of mitochondrial-mediated apoptosis is the release of cytochrome C from the mitochondria into the cytoplasm, a process that is strictly regulated by several membrane-integrated proteins. Predominant among these are the Bcl-2 family of apoptotic regulators. These proteins have been extensively structurally characterized in their inactive, soluble conformations, but perform their primary function after integrating into the outer mitochondrial membrane, having transitioned into an alternative conformation that remains virtually uncharacterized and controversial. We also focus on several integral membrane proteins that reside permanently within the outer mitochondrial membrane (such as MtCH2), for which evidence is accumulating that they may act as receptors for the Bcl-2 proteins, mediating their conformational rearrangements and intermolecular interactions.
KTN(RCK) regulation of K+ channels and transporters
Membrane proteins control the flow in and out of cells of the material necessary for life. One class of these proteins, known as Kef potassium (K+) channels, provide pathogenic bacteria resistance against toxic, antibiotic compounds that would otherwise kill them. These proteins sense certain small molecules called ligands and then open K+-selective channels in the cell membrane as a defensive response. We are studying the mechanisms underlying this critical signaling function, otherwise known as gating, by determining the atomic resolution structures of the proteins at the core of this process. These proteins are purified in large quantities, crystallized, and analyzed through use of intense, highly-focused X-ray radiation. By revealing the 3-dimensional structures of these proteins in complex with both inhibitory and activating ligands, we can see how they sense the presence of these signaling molecules and alter shape, or conformation, in response to that signal.
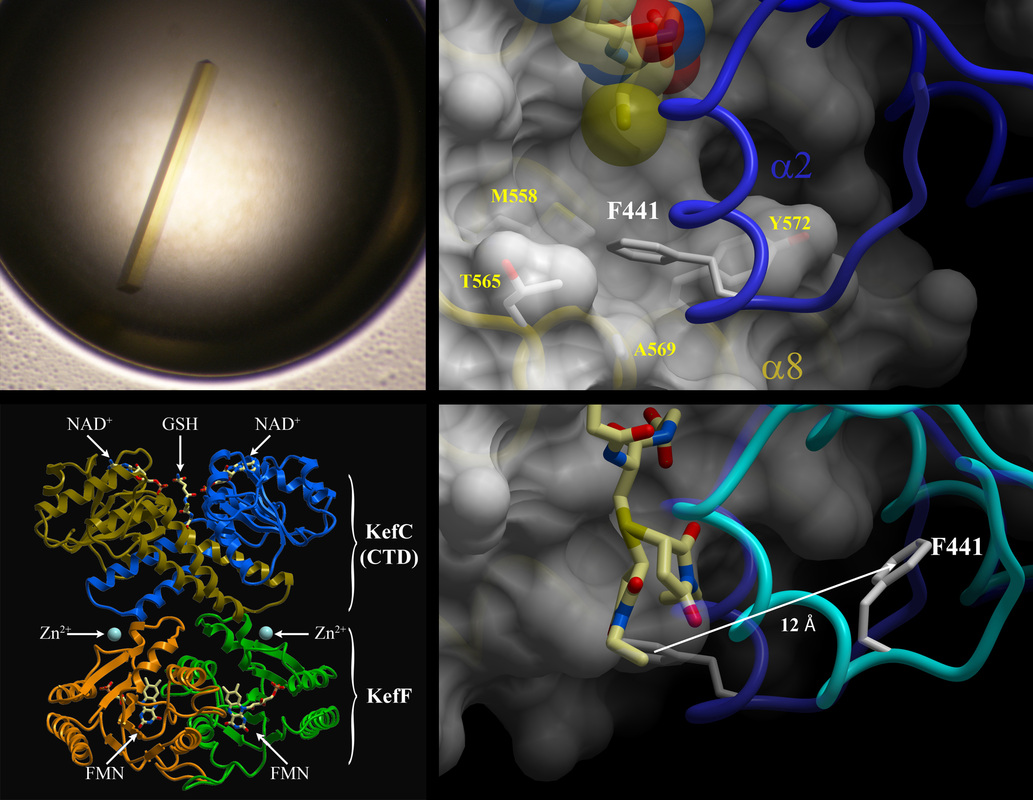
We recently uncovered for the first time a specific ligand-mediated conformational change within a class of regulatory domains called KTN(RCK) that govern K+ flux through several channels, including Kef. The elucidated gating mechanism elegantly explains how the affiliated channel is maintained in a closed, resting state by reduced glutathione (a common cellular molecule involved in chemical detoxification) and yet is activated by glutathione adducts, subtly different molecules formed when a toxin is covalently linked to glutathione. Our results reveal how Kef channels can respond to chemically diverse toxins through a single binding site on the regulatory assembly, providing comprehensive protection to the pathogen against a variety insults.
This breakthrough yields a molecular-level understanding of a key mechanism of transmembrane K+ flux regulation. Given the fundamental role played by these systems in maintaining cellular life, our data should enable rationally designed chemical intervention of this channel’s function, which is applicable to both the development of novel antibiotics and the circumvention of drug resistance in certain pathogenic bacteria such as E. coli, Salmonella and Pseudomonas.
Our laboratory focuses on problems related to carcinogenesis in protein structure, biochemistry and biophysics with an emphasis on membrane proteins. Our understanding of membrane proteins has significantly lagged that of their soluble counterparts due to the numerous difficulties that arise when trying to analyze this class of proteins with conventional biophysical techniques. Arguably, the most acute of these obstacles is in fact simply producing these proteins in sufficient quantities for structural analysis. Most human and, more generally, eukaryotic membrane proteins cannot be expressed in traditional recombinant systems. Our work has led to the discovery of a protein named Mistic that circumvents this primary bottleneck. When fused to a target protein of interest, Mistic acts analogously to a “super-signal sequence,” facilitating the expression of a variety of membrane proteins in their membrane-integrated conformations within the lipid bilayer of recombinant E. coli bacteria. We continue to develop this technology to extend its application to an increasingly diverse array of proteins by surmounting technical difficulties that exist due to steric and topological restrictions imposed by the planar nature of the lipid membrane.
Mitochondrial membrane-associated apoptosis regulators
Mistic is having a major impact on the field of membrane protein structural biology allowing the study of certain scientific problems that were until recently intractable. We are applying this technology to the study of human mitochondrial proteins involved in the regulation of apoptosis. The committing step toward initiation of mitochondrial-mediated apoptosis is the release of cytochrome C from the mitochondria into the cytoplasm, a process that is strictly regulated by several membrane-integrated proteins. Predominant among these are the Bcl-2 family of apoptotic regulators. These proteins have been extensively structurally characterized in their inactive, soluble conformations, but perform their primary function after integrating into the outer mitochondrial membrane, having transitioned into an alternative conformation that remains virtually uncharacterized and controversial. We also focus on several integral membrane proteins that reside permanently within the outer mitochondrial membrane (such as MtCH2), for which evidence is accumulating that they may act as receptors for the Bcl-2 proteins, mediating their conformational rearrangements and intermolecular interactions.
KTN(RCK) regulation of K+ channels and transporters
Membrane proteins control the flow in and out of cells of the material necessary for life. One class of these proteins, known as Kef potassium (K+) channels, provide pathogenic bacteria resistance against toxic, antibiotic compounds that would otherwise kill them. These proteins sense certain small molecules called ligands and then open K+-selective channels in the cell membrane as a defensive response. We are studying the mechanisms underlying this critical signaling function, otherwise known as gating, by determining the atomic resolution structures of the proteins at the core of this process. These proteins are purified in large quantities, crystallized, and analyzed through use of intense, highly-focused X-ray radiation. By revealing the 3-dimensional structures of these proteins in complex with both inhibitory and activating ligands, we can see how they sense the presence of these signaling molecules and alter shape, or conformation, in response to that signal.
We recently uncovered for the first time a specific ligand-mediated conformational change within a class of regulatory domains called KTN(RCK) that govern K+ flux through several channels, including Kef. The elucidated gating mechanism elegantly explains how the affiliated channel is maintained in a closed, resting state by reduced glutathione (a common cellular molecule involved in chemical detoxification) and yet is activated by glutathione adducts, subtly different molecules formed when a toxin is covalently linked to glutathione. Our results reveal how Kef channels can respond to chemically diverse toxins through a single binding site on the regulatory assembly, providing comprehensive protection to the pathogen against a variety insults.
This breakthrough yields a molecular-level understanding of a key mechanism of transmembrane K+ flux regulation. Given the fundamental role played by these systems in maintaining cellular life, our data should enable rationally designed chemical intervention of this channel’s function, which is applicable to both the development of novel antibiotics and the circumvention of drug resistance in certain pathogenic bacteria such as E. coli, Salmonella and Pseudomonas.